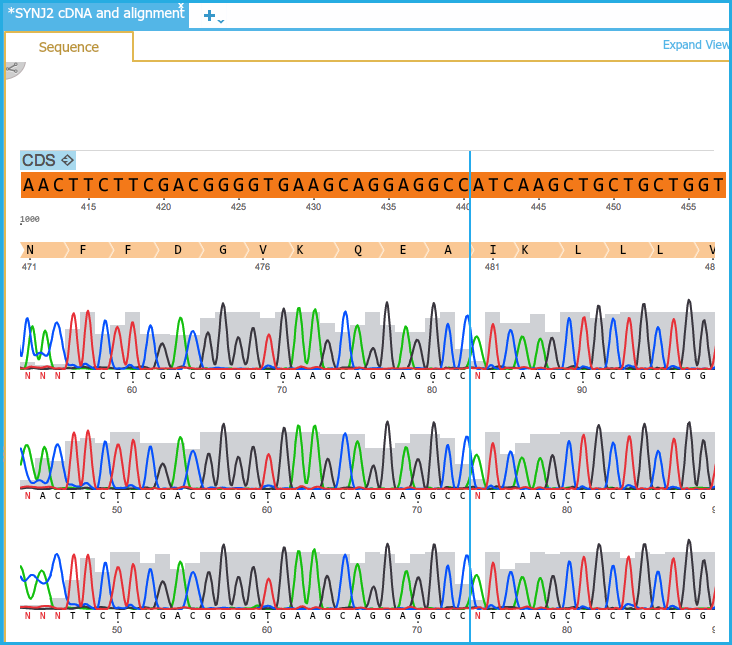
Sangerseq_viewer.sangerseq_viewer provides other useful functions such as generate_consensusseq() and ab1_to_dict() for handling ab1 file. It takes same parameters with sangerseq_viewer command and returns matplotlib.figure object. Turn on 'Show quality values' to see the graph of 'Phred' quality score for each based call. 
If you want to use sanger_seqviewer as python module, please import sangerseq_viewer.sangerseq_viewer and use view_sanger() fucntion. If the option is given, aligned sequences will be output as a Fasta file. f, -output_fasta (optional, default: None) of the mock community was prepared and subjected to preamplification to enrich the target sequences in the mock community dilutions. If the option is given, the values of Sanger sequencing cromatogram will be output as a csv file. c, -output_cromatogram (optional, defualt: None) If output format is pdf, the value is ignored. If True, display bar plot representing Quality value at each nucleotide position. Sangerseq_viewer [-wq (optional, default: True) Sangerseq_viewer -s example_data/puc19_spec_2xu6grna.gb -q example_data/ab1/ -o output/example6.png -l 200 -rs 1700 -re 2100 -dpi 200 Example codeĮxample command 1 sangerseq_viewer -s example_data/puc19_spec_2xu6grna.gb -q example_data/ab1/Spec-2xU6gRNA-1.ab1 -o output/example1.png -dpi 200Įxample command 2 sangerseq_viewer -s example_data/puc19_spec_2xu6grna.gb -q example_data/ab1/Spec-2xU6gRNA-1.ab1 -o output/example2.png -l 200 -dpi 200Įxample command 3 sangerseq_viewer -s example_data/puc19_spec_2xu6grna.gb -q example_data/ab1/Spec-2xU6gRNA-1.ab1 -o output/example3.png-l 200 -rs 1700 -re 2100 -dpi 200Įxample command 4 sangerseq_viewer -s example_data/puc19_spec_2xu6grna.gb -q example_data/ab1/ -o output/example4.png -dpi 200Įxample command 5 sangerseq_viewer -s example_data/puc19_spec_2xu6grna.gb -q example_data/ab1/ -o output/example5.png -l 200 -dpi 200 PREFIX is the executable path of sangerseq_viewer. If you cannnot use sangerseq_viewr command after the installation, please add -prefix=PREFIX option to pip install sangerseq-viewer and try re-installation. You can test Sangerseq_viewer through Google colaboratory. For more information, please refer to their respective documentation. Note: Sangerseq_viewer relies on the packages patchworklib and QUEEN, which provide APIs for managing matplotlib subplots and GenBank files respectively. Sangerseq_viewer provides a solution by allowing you to visualize sequencing results with just a simple command. While commercial GUI software like Snapgene and Geneious Prime offer this functionality, manually processing large amounts of data from Sanger sequencing results can be a tedious and time-consuming task. Sangerseq_viewer is a Python package for visualizing Sanger sequencing results and the corresponding annotated sequence maps automatically.ĭespite being an essential task in DNA sequence construction and editing, there is a lack of open-source software that provides a user-friendly graphical representation of Sanger sequencing results. Must authenticate by NetID.Sangerseq_viewer Installation and User Manual Geneious Prime- UArizona License for UArizona users Editing is not supported with the free version Snap Gene Viewer is a free download to view traces. It is a trace viewer for the PC platform (all Windows OS™s are supported). If you need assistance with large scale sequencing data management, please feel free to contact us for assistance or recommendations.Ĭhromas is freely available over the web, and offers many options for working with. These programs are for viewing individual sequence files. Support for the addition of new plasmid features and new plasmids has been added, giving users greater control over the content of PlasMapper and a richly. There are several free viewers that are available to look at the electropherograms. PlasMapper 3.0 has several interactive text editors/viewers that allow users to select and view plasmid regions, insert genes, modify restriction sites or perform codon optimization. Please see the notes at the bottom of this page on how to configure your computer's file associations so that the files will open when you click on them. 
As you might expect, these viewers are platform specific. The files that our automated sequencers generate require specialized software in order to be viewed.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
